About the CoMSES Model Library more info
Our mission is to help computational modelers develop, document, and share their computational models in accordance with community standards and good open science and software engineering practices. Model authors can publish their model source code in the Computational Model Library with narrative documentation as well as metadata that supports open science and emerging norms that facilitate software citation, computational reproducibility / frictionless reuse, and interoperability. Model authors can also request private peer review of their computational models. Models that pass peer review receive a DOI once published.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and feel free to contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 6 of 46 results for "Christine Schnor" clear search
Peer reviewed FISHCODE - FIsheries Simulation with Human COmplex DEcision-making
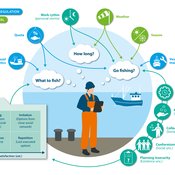
Birgit Müller Gunnar Dressler Jonas Letschert Christian Möllmann Vanessa Stelzenmüller | Published Monday, August 05, 2024FIsheries Simulation with Human COmplex DEcision-making (FISHCODE) is an agent-based model to depict and analyze current and future spatio-temporal dynamics of three German fishing fleets in the southern North Sea. Every agent (fishing vessel) makes daily decisions about if, what, and how long to fish. Weather, fuel and fish prices, as well as the actions of their colleagues influence agents’ decisions. To combine behavioral theories and enable agents to make dynamic decision, we implemented the Consumat approach, a framework in which agents’ decisions vary in complexity and social engagement depending on their satisfaction and uncertainty. Every agent has three satisfactions and two uncertainties representing different behavioral aspects, i.e. habitual behavior, profit maximization, competition, conformism, and planning insecurity. Availability of extensive information on fishing trips allowed us to parameterize many model parameters directly from data, while others were calibrated using pattern oriented modelling. Model validation showed that spatial and temporal aggregated ABM outputs were in realistic ranges when compared to observed data. Our ABM hence represents a tool to assess the impact of the ever growing challenges to North Sea fisheries and provides insight into fisher behavior beyond profit maximization.
Benchmark for DMASON
Andreu Moreno Vendrell Josep Jorba Eduardo César Anna Sikora Cristina Peralta Quesada | Published Friday, November 22, 2024Agent-based modeling and simulation (ABMS) is a class of computational models for simulating the actions and interactions of autonomous agents with the goal of assessing their effects on a system as a whole. Several frameworks for generating parallel ABMS applications have been developed taking advantage of their common characteristics, but there is a lack of a general benchmark for comparing the performance of generated applications. We propose and design a benchmark that takes into consideration the most common characteristics of this type of applications and includes parameters for influencing their relevant performance aspects. We provide an initial implementation of the benchmark for DMASON parallel ABMS platform, and we use it for comparing the applications generated by these platforms.
00b SimEvo_V5.08 NetLogo
Garvin Boyle | Published Saturday, October 05, 2019In 1985 Dr Michael Palmiter, a high school teacher, first built a very innovative agent-based model called “Simulated Evolution” which he used for teaching the dynamics of evolution. In his model, students can see the visual effects of evolution as it proceeds right in front of their eyes. Using his schema, small linear changes in the agent’s genotype have an exponential effect on the agent’s phenotype. Natural selection therefore happens quickly and effectively. I have used his approach to managing the evolution of competing agents in a variety of models that I have used to study the fundamental dynamics of sustainable economic systems. For example, here is a brief list of some of my models that use “Palmiter Genes”:
- ModEco - Palmiter genes are used to encode negotiation strategies for setting prices;
- PSoup - Palmiter genes are used to control both motion and metabolic evolution;
- TpLab - Palmiter genes are used to study the evolution of belief systems;
- EffLab - Palmiter genes are used to study Jevon’s Paradox, EROI and other things.
…
Gradient Descent Simulation
Ilyes Azouani | Published Wednesday, March 18, 2026This model visualizes gradient descent optimization - the fundamental algorithm used to train neural networks and other machine learning models. Agents represent different optimization algorithms searching for the minimum of a loss landscape (the “error surface” that ML models try to minimize during training).
The model demonstrates how different optimizer types (SGD, Momentum with different parameters) behave on various loss landscapes, from simple bowls to the notoriously difficult Rosenbrock “banana valley” function. This helps build intuition about why certain optimization algorithms work better than others for different problem geometries.
HOW IT WORKS
…
An agent-based model exploring the biodiversity-agriculture-nutrition dynamics in Eastern Madagascar resulting from alternative land uses
Romain Clercq-Roques Jessica Williams Katja Perez Guzman Marta Kozicka Kristine Belesova Zaid Chalabi | Published Wednesday, July 23, 2025This agent-based model simulates the interactions between smallholder farming households, land-use dynamics, and ecosystem services in a rural landscape of Eastern Madagascar. It explores how alternative agricultural practices —shifting agriculture, rice cultivation, and agroforestry—combined with varying levels of forest protection, influence food production, food security, dietary diversity, and forest biodiversity over time. The landscape is represented as a grid of spatially explicit patches characterized by land use, ecological attributes, and regeneration dynamics. Agents make yearly decisions on land management based on demographic pressures, agricultural returns, and institutional constraints. Crop yields are affected by stochastic biotic and abiotic disruptions, modulated by local ecosystem regulation functions. The model additionally represents foraging as a secondary food source and pressure on biodiversity. The model supports the analysis of long-term trade-offs between agricultural productivity, human nutrition, and conservation under different policy and land-use scenarios.
Narragansett Bay (RI) Recreational Fishery ABM
Tyler Pavlowich Anne Innes-Gold Margaret Heinichen M. Conor McManus Jason McNamee Jeremy Collie Austin Humphries | Published Monday, June 21, 2021This model is based on the Narragansett Bay, RI recreational fishery. The two types of agents are piscivorous fish and fishers (shore and boat fishers are separate “breeds”). Each time step represents one week. Open season is weeks 1-26, assuming fishing occurs during half the year. At each weekly time step, fish agents grow, reproduce, and die. Fisher agents decide whether or not to fish based on their current satisfaction level, and those that do go fishing attempt to catch a fish. If they are successful, they decide whether to keep or release the fish. In our publication, this model was linked to an Ecopath with Ecosim food web model where the commercial harvest of forage fish affected the biomass of piscivorous fish - which then became the starting number of piscivorous fish for this ABM. The number of fish caught in a season of this ABM was converted to a fishing pressure and input back into the food web model.
Displaying 6 of 46 results for "Christine Schnor" clear search